My current project is re-identifying individual manta rays (Mobula alfredi and Mobula birostris) by their distinct belly patterns … er… ventral markings.
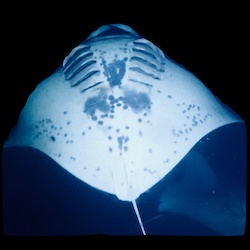
Every night at two spots on the Big Island of Hawai’i where I live, dive boats shine bright lights that attract plankton. Most nights, the plankton in turn attract multiple manta rays to feed. The mantas, most of which are between 3 and 4 meters in wingspan, form feeding trains, barrel-roll in the plankton columns, and come within inches of the divers and snorkelers. I’ve dived all over the world, and I think the Kona manta dive is among the very best large-animal dives in all the world.
The dives have a huge economic impact: every year, more than 100,000 people snorkel or dive with the mantas in Kailua Kona and generate well over $10M in revenue. (Those numbers are from 2016 and obviously the pandemic slowed things down. More recently, the growth in manta snorkeling has triggered some much-needed regulatory review.)
Re-identifying animals is a common problem among field and population biologists. Generally, new photos are matched against a photo catalog by a freakishly-accurate grad student or longtime researcher (every species has their own freakishly-accurate identifier!). The work is tedious and often backlogged.
For animals with prominent markings (like mantas), there are a number of good pattern-recognition algorithms that are implemented in OpenCV, the broadly-used computer vision library: SIFT, SURF, and ORB. Mantas are almost laughably amenable to geometry-based algorithms: it’s only one side that has distinctive markings, the body of the manta is about as flat a surface as you’ll find in the animal kingdom, and the markings are generally high-contrast black-and-white. The Manta Pacific Research Foundation, with whom I’m working, currently uses SIFT for re-identification.
In recent years, deep-learning techniques for individual re-identification have been successfully applied to a number of species: happywhale in particular, has catalogs of more than two dozen cetacean species. Deep learning has been successfully applied to mantas, too, so there’s every reason to believe that I should be able to replicate their success.
While mantas are a happy subject for me, since I literally see the dive boats heading out to the manta ray spots every night, I am more generally interested in working towards a “bring your own species” re-identification system. The techniques used for re-identification using deep learning are now well-established in the ML world. Computer Science grad students are unlikely to devote themselves to fine-tuning known algorithms and creating an efficient end-to-end pipeline and Biology grad students may tend to write overly-specific code. Me, on the other hand? I’ve always been a “technology transfer” coder, not a researcher, and have no problem working downstream of the latest-and-greatest. Meanwhile, I have a lot of experience operationalizing and scaling software systems.
So, I hope, this project will be a good combination of my interests and talents, in service to environmental causes.